Example 2: Using the modeling interface
This example demonstrates how to manually construct a graph instance using the Model class. We build a small network from scratch and solve a Critical Node Problem with variable population sizing.
Problem description
We construct a graph with 20 nodes and 22 edges, then solve a CNP instance where we remove 3 nodes to minimize the pairwise connectivity of the remaining graph. This example also demonstrates the Variable Population Memetic Search (VPMS) algorithm.
Building the graph
First, we initialize an empty model and add nodes:
from pycnp import Model
from pycnp.MemeticSearch import MemeticSearchParams, VariablePopulationParams
from pycnp.stop import NoImprovement
from pycnp.visualization import visualize_graph
# 1. Initialize the model
model = Model()
# 2. Add 20 nodes (0-19)
for i in range(20):
model.add_node(i)
Adding edges defines the network structure. Each edge connects two nodes in an undirected graph:
# 3. Add edges to define the graph structure
edges = [
(0, 4), (1, 4), (2, 4), (2, 6), (3, 4), (4, 9), (5, 9),
(6, 9), (7, 9), (7, 11), (8, 9), (9, 14), (10, 14), (11, 14),
(12, 14), (12, 16), (13, 14), (14, 19), (15, 19), (16, 19),
(17, 19), (18, 19)
]
for u, v in edges:
model.add_edge(u, v)
Configuring problem parameters
We define the problem type, budget (number of nodes to remove), and stopping criterion:
# 4. Configure problem parameters
problem_type = "CNP"
budget = 3
seed = 6
stopping_criterion = NoImprovement(20)
Configuring the memetic algorithm
The MemeticSearchParams class configures the memetic search algorithm. We use:
search: Local search strategy (
"CHNS"- Component-Based Hybrid Neighborhood Search)crossover: Crossover operator (
"RSC"- Reduce-Solve-Combine)reduce_params: Parameters for the RSC crossover (search strategy and beta value)
is_pop_variable: Enable variable population sizing
initial_pop_size: Initial population size
# 5. Configure memetic search with variable population
memetic_params = MemeticSearchParams(
search="CHNS",
crossover="RSC",
reduce_params={"search": "CHNS", "beta": 0.8},
is_pop_variable=True,
initial_pop_size=5,
)
Variable population sizing adapts the population size during search based on convergence behavior. The VariablePopulationParams class configures this:
pop_params = VariablePopulationParams(
max_pop_size=20, # Maximum population size
increase_pop_size=3, # Individuals to add when expanding
max_idle_gens=20 # Generations without improvement before expansion
)
Solving the problem
We solve the instance using the configured parameters:
# 6. Solve the instance via VPMS (Variable Population Memetic Search)
result = model.solve(
problem_type,
budget,
stopping_criterion,
seed,
memetic_params,
pop_params,
display=True,
)
The display=True parameter enables progress output during solving.
Results
After solving, we can access the solution:
print(f"Removed nodes: {result.best_solution}")
print(f"Remaining connectivity: {result.best_obj_value}")
print(f"Number of iterations: {result.num_iterations}")
print(f"Runtime: {result.runtime:.2f} seconds")
You obtain output similar to:
Removed nodes: {9, 19, 14}
Remaining connectivity: 17
Number of iterations: 20
Runtime: 1.28 seconds
Visualizing the result
PyCNP provides an interactive graph visualization using the pyvis library. The visualize_graph() function generates an HTML file showing the original graph with critical nodes highlighted:
# 7. Visualize the result
visualize_graph(
problem_data=model.problem_data,
critical_nodes_set=result.best_solution,
)
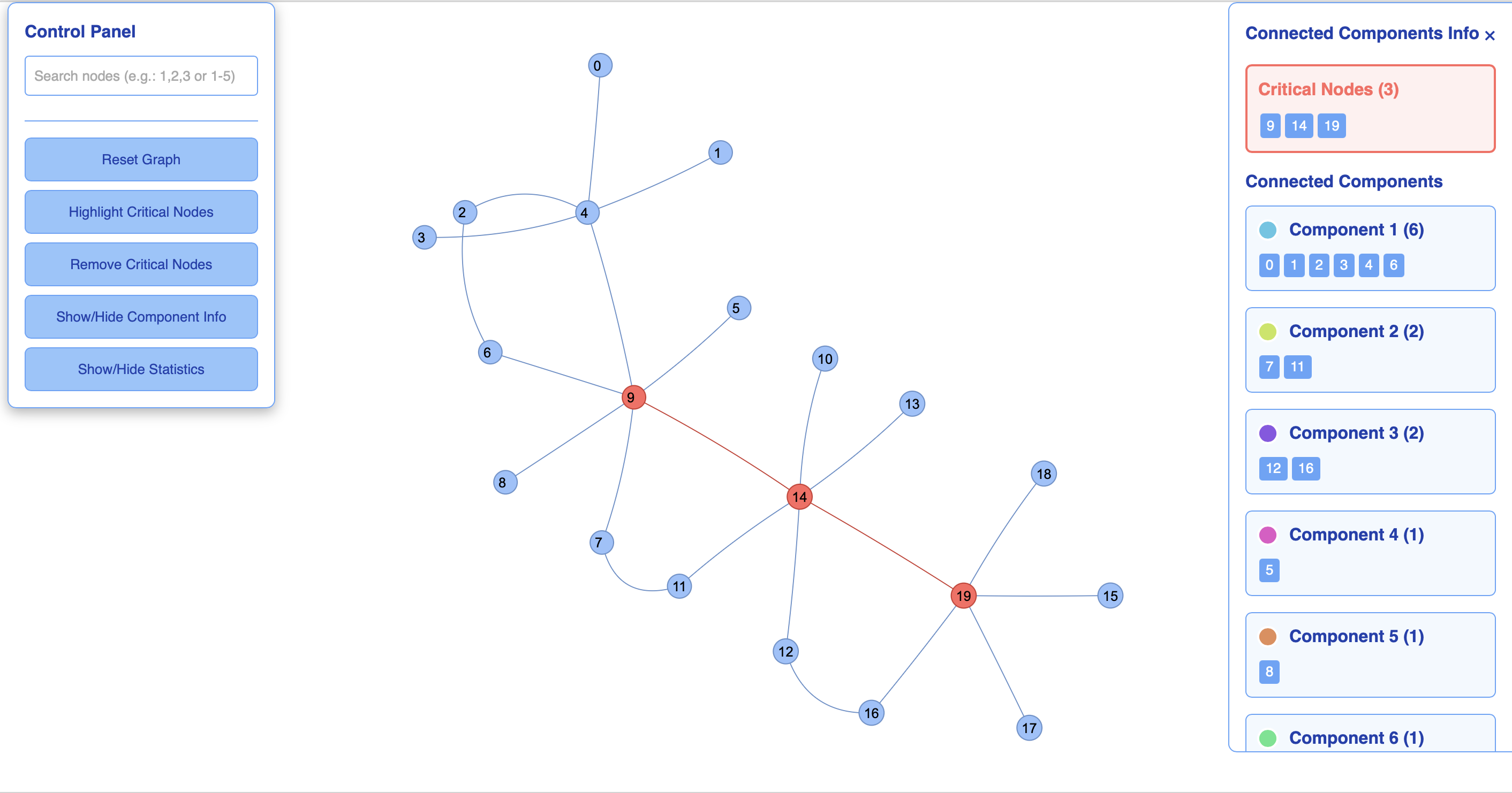
This generates an interactive HTML file that opens automatically in your browser. Critical nodes are displayed in a different color, and you can interact with the visualization.
Understanding the output
best_solution: Set of node IDs removed from the graph
best_obj_value: Pairwise connectivity of the remaining graph
num_iterations: Number of algorithm iterations performed
runtime: Total execution time in seconds
Note
The variable population mechanism adapts the population size dynamically:
- If no improvement is observed for max_idle_gens, the population expands by increase_pop_size
- The population is capped at max_pop_size